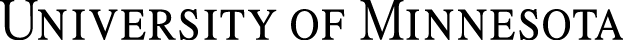73 Molecules Talk: Selecting Molecular Communication and Complexity
In our complex human society, we define communication by its specificity. Without a careful choice of words, our speech would be at best, a source of magnificent misunderstanding, or just plain babel! What does this mean for prebiotic chemistries?
In terms of prebiotic chemical evolution, selection by definition would have favored protective accumulation of longer-lived molecular aggregates. Over time, the same selective imperatives would create webs of such aggregates, increasing the range and specificity of molecular interactions in a challenging environment. If this were to have occurred in an enclosed proto-cellular space, it should have resulted in a primitive molecular communication and a growing complexity (another property of life!). In fact, alof the properties of life must have accompanied the achievement of more and more complex intermolecular communication. Simply put, a prebiotic (or for that matter a cellular) genetic change that alters the rate of one catalytic reaction (if not destructive) will drive the selection of changes in components of other, interconnected metabolic chemistries. If molecular communication required the evolution of catalytic specificity, then the final elaboration of complexity and order as a property of life further requires the selection of mechanisms of regulation and coordination.
Intermolecular Communication Leads to an Early Establishment of Essential Interconnected Chemistries
Earlier, we suggested that inorganic catalyst precursors to biological enzymes were probably minerals embedded in clay or other substrata, providing surfaces that would naturally aggregate organic molecules and catalyze repetitive reactions. Either the initial objects of prebiotic selection included external stable monomers and polymers, outside or as seems more likely, inside proto-cells. Later, selection would have favored polymers that enhanced growth and reproduction of successful aggregates. These polymers were likely those that catalyzed their own synthesis, perhaps collaborating with inorganic catalytic minerals. The result would be the elaboration of a web of interconnected chemical reactions between molecules with high affinity for each other, thereby increasing the specificity of those reactions. In the context of life origins and evolution, co-catalysis describes the activities of these interconnected metabolic reactions.
As noted, high-affinity interactions are inherently protective. During prebiotic chemical/metabolic evolution, protected stable molecular assemblies would be targets of selection. Continuing co-evolution of catalysts, substrates and co-catalytic reaction sets would lead to more and more sophisticated molecular communication. Once established, efficient biochemical reaction sets would be constrained against significant evolutionary change. Any change (mutation) that threatened this efficiency would mean the end of a prebiotic chemical (or for that matter, cell) lineage! This explains why we find common pathways for energy-generation (e.g., autotrophic and fermentative), reproduction (replication), and information storage and retrieval (DNA, RNA, protein synthesis) in all of LUCA’s descendants.
Sophisticated, effective communication requires coordination. In fact, effective communication is defined by coordination, the capacity to make chemical decisions. Selection of molecular aggregates that sequestered metabolic reactions behind a semipermeable membrane ensure that only certain molecules communicate with each other. This sequestration is likely to have occurred repeatedly during chemical evolution, beginning with the synthesis of larger, polymeric molecules and possibly, an aggregation of primitive lipoidal molecules. We can think of increasingly effective catalysis in an enclosed environment as a conversation mediated by good speakers! Coordination is a property that likely co-evolved with life itself!
Origins of Coordination
Let’s look some possible structures churning around in the prebiotic chemistry set that might have self-assembled, or sequestered compatible chemistries of life. Along with the alkaline vent biofilm compartment, coacervates, proteinoid microspheres and liposomes have been considered as possible progenitors of biological membranes. Each can be made in the laboratory. They are demonstrably semipermeable, and in some cases can even replicate! Micrographs and the production of coacervates, proteinoid microspheres and liposomes are shown below (Figure 1).

Oparin had proposed that the action of sunlight in the absence of oxygen could cause ionized, oppositely charged organic molecules (e,g, amino acids, carbohydrates, etc.) to form droplets from organic molecules in his primordial soup. These coacervates were actually produced in 1932, visualized by microscopy and demonstrated to be a semi-permeable compartment. They even behaved as if they were able to grow and reproduce (also as Oparin originally suggested they might).
In the 1950s, Sidney Fox produced proteinoid microspheres from short peptides that formed spontaneously from aqueous amino acid solutions heated to dryness (not unlike what happens in the tidal pool scenario of polymer formation from organic monomers). These can be seen by light and electron microscopy.
While liposomes are easily made in a laboratory, it isn’t clear that they existed on a pre-biotic earth. Nevertheless, cell membranes must have had acquired their phospholipid bilayer structure by the time of LUCA since we all have them! Prior to LUCA, chemical rearrangenments must have occurred to enable incorporation of a phospholipid bilayer into whatever starting boundary life started with.
We have already considered the biofilm proposed for cellular origins in an alkaline vent. The formation of such biofilms would have separated acidic ocean protons from the interior of such protocells, creating a proton gradient. Such a gradient could have driven the early evolution of chemiososis as a means to create chemical energy, complete with the eventual selection of ATP synthases and the enzymes of proton transport, again because all cells descendent from LUCA’s posess these biochemistries. Of course, proteinoid microspheres, coacervates, biofilm-based ‘membranes and liposomes are not alive, and are therefore not cells. But one or another of them must have been where the enhanced coordination of molecular communication required for life began.
Watch this video to learn about Protected Molecular Communication: Semipermeable Membranes.
An important take-home message here is that whatever the original structure of the first cells, they arose soon after the organic chemical prerequisites of life began to acquire familiar metabolic functions. We need to see chemical and structural progress to cellularity as concurrent metabolic evolutionary events. At some point, selection of sequestered biochemistries led to protocells, then to the first cells, each with all of the properties of life.
Finally, selection of highly specific communication between cellular molecules allowed cells themselves to talk to one another, engage in group activities, and eventually join together to form multicellular organisms. Multicellularity is of course a characteristic of many if not most eukaryotes. But watch a great TED Talk on bacterial intercellular communication by Dr. Bonnie Bassler at Intercellular Communication in Bacteria.
Origins of Information Storage and Retrieval in an RNA World
Let us accept for now that molecular communication began concurrently with the packaging of interconnected co-catalytic sets into semipermeable structures. Then the most ‘fit’ of these structures were selected for efficient coordination of meaningful, timely chemical messages. Ultimately, coordination requires information processing, storage and retrieval, something we recognize in Francis Crick’s Central Dogma of information flow from DNA to RNA to protein. Cells and organisms do coordination quite well, but what do its beginnings look like? The answer may lie in the pre-biotic RNA world we discussed earlier. The Central Dogma, modified to account for reverse transcription and the behavior of retroviruses, is shown below (Figure 2).
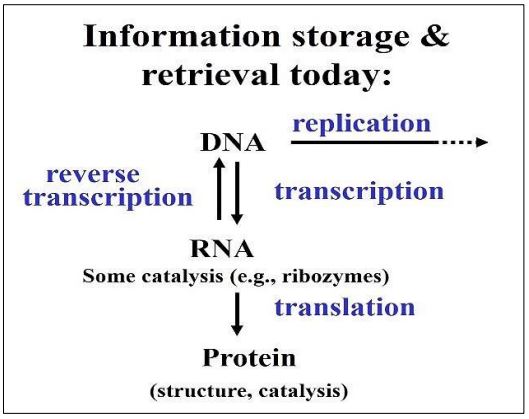
We do not really know how cells came to rely on DNA to store, pass on and mobilize genetic information, but we have presented reasons to believe that the first replicating nucleic acid was RNA, creating an RNA world. Here is the evidence that leads us to this conclusion.
- Based on the stem-and-loop and other structures that form when RNA molecules undergo internal H-bonding, we know that RNAs can take on varied and intricate shapes.
- Diverse conformations are consistent with the evolution of specificity in the interaction of RNAs with themselves and/or with other molecules in the prebiotic environment.
- RNAs, either alone as autocatalysts (for example, self-splicing mRNAs) or in catalytic ribonucleoprotein complexes (e.g., ribosomes) exist in cells today.
- Some of these RNAs (specifically rRNAs), have a long phylogenetic heritage, shared by cells in all three domains of life.
The propensity of single stranded RNA molecules to fold based on internal H-bonding can lead to those diverse three-dimensional shapes (tertiary structure). These structures could have interacted with other molecules in a environment. Because they could be replicated according to different prebiotic scenarios, the same RNAs could also pass on simple genetic information contained in their base sequences. The combination of informational and catalytic properties in a single molecule is illustrated below (Figure 3).
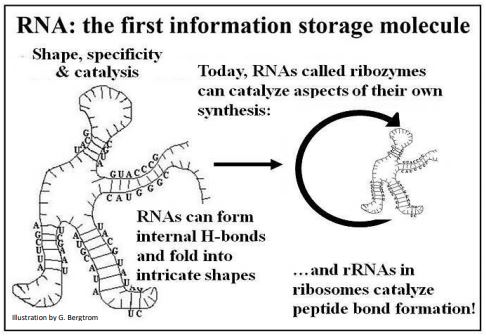
The capacity of RNAs as catalysts and warehouses of genetic information speaks to an efficient candidate for the first dual or multi-purpose polymer, a property that is not known and cannot be demonstrated for DNA. Read more about the proposed ‘RNA worlds’ in which life may have begun in Cech TR (2012) [The RNA Worlds in Context. In Cold Spring Harbor Perspectives in Biology (Cold Spring Harbor, NY: Cold Spring Harbor press) 4(7):a006742e].
Watch this video to learn about Self-Replication: Information, Communication, and Coordination.
What might RNA catalysis beyond self-replication have looked like in simpler times? Consider the interaction between a two hypothetical RNAs and different hypothetical amino acids bound to each, shown below (Figure 4).
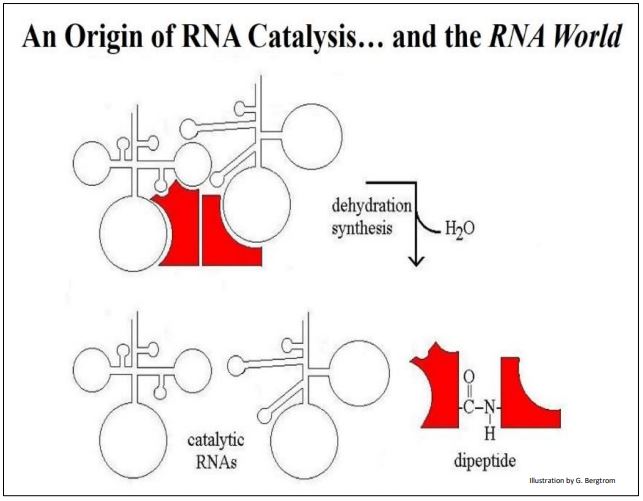
The binding of each RNA to its amino acid would be a high affinity, specific interaction based on charge and shape complementarity. Likewise, the two RNAs seen in the illustration must have a high affinity for each other, also based on chemical and physical complementarities. One can even envision some strong H-bonding between bases in the two RNAs that might displace intra-strand H-bonding (not shown here). The result is that the two amino acids are brought together in a way that catalyzes peptide bond formation. This will require an input of free energy (recall that peptide bond is one of the most energy intensive reaction in cells). For now, assume a chemical energy source and let us focus on the specificities required for RNA catalytic activity.
We know now that tRNAs the intermediaries between nucleic acids and polypeptide synthesis. So it’s fair to ask how the kind of activity illustrated above could have led to the tRNA-amino acid interactions we see today. There is no obvious binding chemistry between today’s amino acids and RNAs, but there may be a less obvious legacy of the proposed bindings. This has to do with the fact that the genetic code is universal, which means that any structural relationship between RNA and amino acids must have been selected early (at the start!) of cellular life on earth. Here is the argument.
- The code is indeed universal (or nearly so)
- There is a correlation between the chemical properties of amino acids and their codons, for example:
- Charged (polar) amino acids are encoded by triplet codons with more G (guanine) bases.
- Codons for uncharged amino acids more often contain a middle U (uracil) than any other base.
These correlations would mean that an early binding of amino acids to specifically folded RNAs was replaced in evolution by enzyme-catalyzed covalent attachment of an amino acid to a ‘correct’ tRNA, such as we see today.
What forces might have selected separation of the combined template/informational functions from most of the catalytic activities of RNAs? Perhaps it was the selection of the greater diversity of structure (i.e., shape) that folded polypeptides can achieve, compared to folded RNAs. After all, polypeptides are strings of 20 different amino acids compared to the four bases that make up nucleic acids. This potential for molecular diversity would in turn accelerate the pace of chemical (and ultimately cellular) evolution. A scenario for the transition from earlier self-replicating RNA events to the translation of proteins from mRNAs is illustrated here (Figure 5).
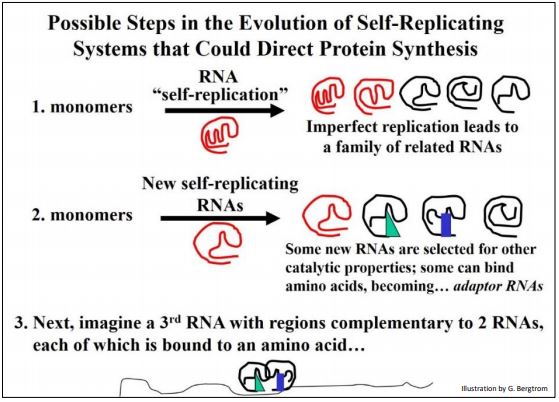
Adaptor RNAs in the illustration will become tRNAs. The novel, relatively unfolded RNA depicts a presumptive mRNA. Thus, even before the entry of DNA into our RNA world, it is possible to imagine the selection of the defining features of the genetic code and mechanism of translation (protein synthesis) that characterizes all life on the planet. Next, we consider “best-speculations” of how RNA-based information storage and catalytic chemistries might have made the evolutionary transition to DNA-based information storage and predominantly protein based enzyme catalysis.
From Ribosomes to Enzymes; From RNA to DNA
The term co-catalysis could very well describe biochemical reactions in which a catalyst accelerates a chemical reaction whose product feeds back in some way on its own synthesis. We saw this in action when we discussed allosteric enzyme regulation and the control of biochemical pathways. Catalytic feedback loops must have been significant events in the evolution of the intermolecular communication and metabolic coordination required for life.
Here we will consider some scenarios for the transition from an RNA world to something more recognizable as today’s nucleic acid information storage and proteinbased catalytic metabolism.
1. Ribozymes Branch Out: Replication, Transcription, and Translation
If RNAs catalyzed their own replication, it may have resembled the autocatalytic replication of AATE. At the same time, some RNAs may also have attracted amino acids to their surfaces and catalyzed peptide bond formation, as already described. Shapely prebiotic RNAs may therefore have catalyzed synthesized peptides, some of which would eventually take over catalysis of RNA synthesis! The scenario is summarized below (Figure 6).
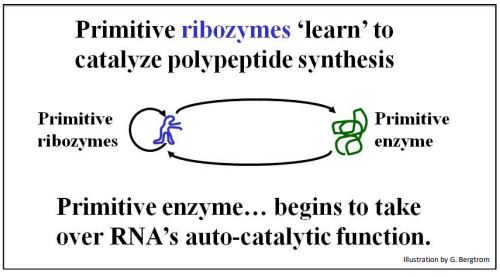
Watch this video to learn about Information Storage and Retrieval in an RNA World.
Selection favoring the synthesis of short oligopeptides and polypeptides is consistent with a catalytic diversification that led to the dominance of protein catalysts, i.e., enzymes. The primitive enzyme shown here must have been selected because at first, it assisted the autocatalytic replication of the RNA itself! Over time, the enzyme would evolve along with the RNA. This co-evolution then eventually replaced autocatalytic RNA replication with the enzyme-catalyzed RNA synthesis we recognize as transcription today. In this scenario, self-splicing premRNAs and ribozymes are surviving remnants of an RNA world!
Let’s turn now to some ideas about how an RNA world could make the transition to the DNA-RNA-protein world we have today
2. Transfer of Information Storage from RNA to DNA
The transfer of function from RNA to DNA is by no means a settled issue among students of life origins and early evolution. A best guess is that the elaboration of protein enzymes begun in the RNA world would lead to reverse transcriptase-like enzymes that copied RNA information into DNA molecules. DNA information may have been selected because DNA is chemically more stable than RNA. The basic transfer scenario is illustrated below (Figure 7).
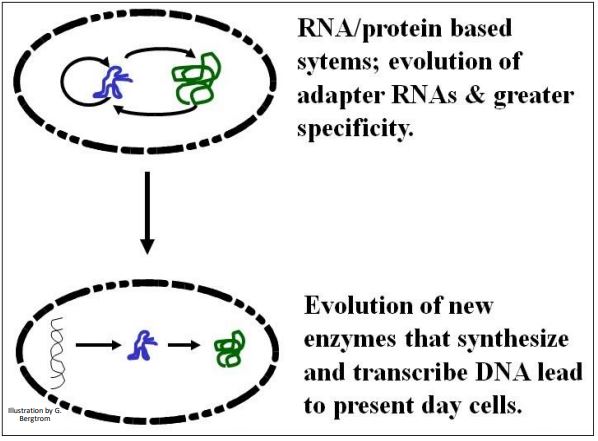
All cells alive today store information in DNA (only some viruses have an RNA genome). Therefore, transition to the use of DNA as an information molecule must have preceded the origin of life. At least, it must have occurred in the cells from which the LUCA arose. Details of this key change involve evolutionary steps yet to be worked out to everyone’s satisfaction!
Watch this video to learn about the Transition from an RNA World to a DNA World.
The Evolution of Biochemical Pathways
The tale of the evolution of enzymes from ribozymes and of informational DNA from RNA, and the other metabolic chemistries behind prebiotic semipermeable boundaries is ongoing in cells today. Undoubtedly, early cellular metabolism involved only reactions crucial to life, catalyzed by a limited number of enzymes. But, if evolution inexorably trends towards greater complexity of molecular communication and coordination, in other words, towards increasingly refined regulation of metabolism, how did the repertoire of enzymes get larger, and how did biochemical pathways become more elaborate? We answered the first question elsewhere, when we discussed gene duplication (e.g., by unequal crossing over). The duplicate genes encoding the same enzyme provided the raw material for new enzymes and new enzymatic functions.
Whether in cells or in prebiotic structures, we can hypothesize how a new chemical reaction could evolve. For example, assume that a cell acquires molecule D required for an essential function, from an external, environmental source. What happens if levels of D in the environment become limiting? Clearly, cells would die without enough D. That is, unless cells that already have a duplicated, redundant gene that has mutated and now encodes an enzyme with the ability to make D in the cell. Such a cell might have existed with other cells without the mutation, but a D-limited environment would select the mutant cell for survival and reproduction. Imagine the scenario illustrated below (Figure 8).
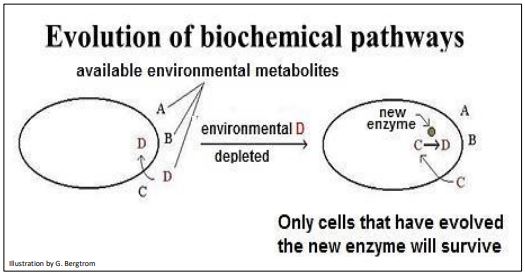
Watch this video to learn about the Origins and Evolution of Biochemical Pathways.
In a similar scenario, a mutation in a duplicated gene could result in a novel enzyme activity that can convert some molecule (e.g., C or D) in the cell into a new molecular product. If the new enzyme and molecular product do not kill or debilitate the cell, the cell might survive to be selected by some future exigency.
This chapter by Gerald Bergtrom is licensed CC BY 4.0.