102 Viral Evolution, Morphology, and Classification
By the end of this section, you will be able to do the following:
- Describe how viruses were first discovered and how they are detected
- Discuss three hypotheses about how viruses evolved
- Describe the general structure of a virus
- Recognize the basic shapes of viruses
- Understand past and emerging classification systems for viruses
- Describe the basis for the Baltimore classification system
Viruses are diverse entities: They vary in structure, methods of replication, and the hosts they infect. Nearly all forms of life—from prokaryotic bacteria and archaeans, to eukaryotes such as plants, animals, and fungi—have viruses that infect them. While most biological diversity can be understood through evolutionary history (such as how species have adapted to changing environmental conditions and how different species are related to one another through common descent), much about virus origins and evolution remains unknown.
Discovery and Detection
Viruses were first discovered after the development of a porcelain filter—the Chamberland-Pasteur filter—that could remove all bacteria visible in the microscope from any liquid sample. In 1886, Adolph Meyer demonstrated that a disease of tobacco plants—tobacco mosaic disease—could be transferred from a diseased plant to a healthy one via liquid plant extracts. In 1892, Dmitri Ivanowski showed that this disease could be transmitted in this way even after the Chamberland-Pasteur filter had removed all viable bacteria from the extract. Still, it was many years before it was proved that these “filterable” infectious agents were not simply very small bacteria but were a new type of very small, disease-causing particle.
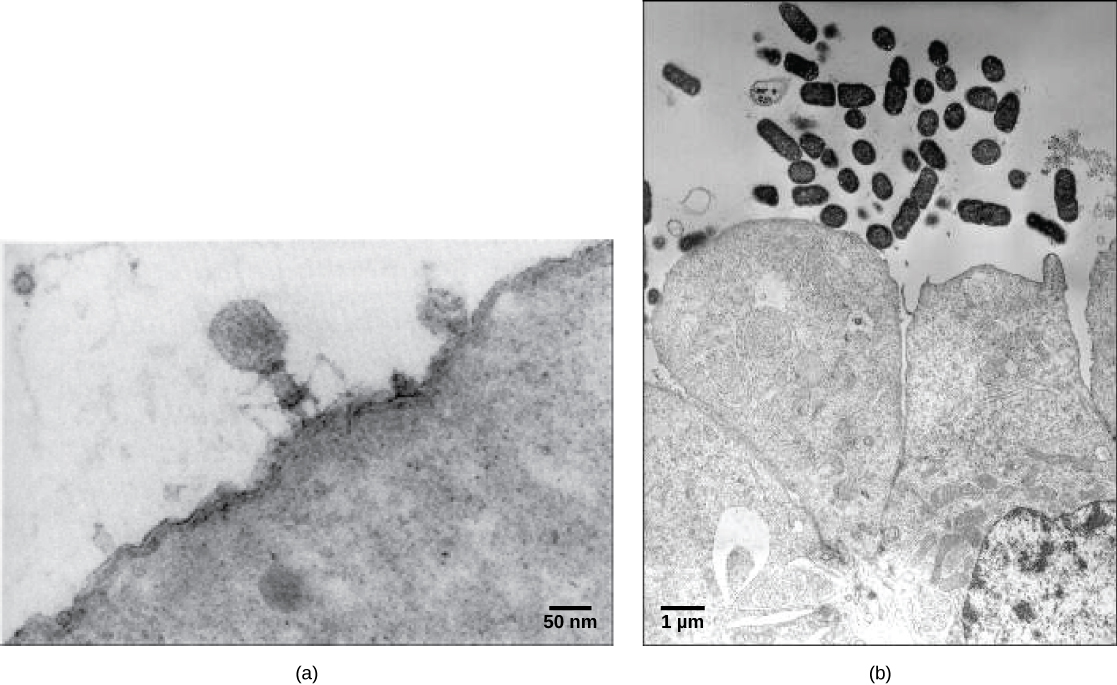
Most virions, or single virus particles, are very small, about 20 to 250 nanometers in diameter. However, some recently discovered viruses from amoebae range up to 1000 nm in diameter. With the exception of large virions, like the poxvirus and other large DNA viruses, viruses cannot be seen with a light microscope. It was not until the development of the electron microscope in the late 1930s that scientists got their first good view of the structure of the tobacco mosaic virus (TMV) (Viruses: Introduction: Figure 1), discussed in the previous chapter, and other viruses (Figure 1, below). The surface structure of virions can be observed by both scanning and transmission electron microscopy, whereas the internal structures of the virus can only be observed in images from a transmission electron microscope. The use of electron microscopy and other technologies has allowed for the discovery of many viruses of all types of living organisms.
Evolution of Viruses
Although biologists have a significant amount of knowledge about how present-day viruses mutate and adapt, much less is known about how viruses originated in the first place. When exploring the evolutionary history of most organisms, scientists can look at fossil records and similar historic evidence. However, viruses do not fossilize, as far as we know, so researchers must extrapolate from investigations of how today’s viruses evolve and by using biochemical and genetic information to create speculative virus histories.
Most scholars agree that viruses don’t have a single common ancestor, nor is there a single reasonable hypothesis about virus origins. There are current evolutionary scenarios that may explain the origin of viruses. One such hypothesis, the “devolution” or the regressive hypothesis, suggests that viruses evolved from free-living cells, or from intracellular prokaryotic parasites. However, many components of how this process might have occurred remain a mystery. A second hypothesis, the escapist or the progressive hypothesis, suggests that viruses originated from RNA and DNA molecules that escaped from a host cell. A third hypothesis, the self-replicating hypothesis, suggests that viruses may have originated from self-replicating entities similar to transposons or other mobile genetic elements. In all cases, viruses are probably continuing to evolve along with the cells on which they rely on as hosts.
As technology advances, scientists may develop and refine additional hypotheses to explain the origins of viruses. The emerging field called virus molecular systematics attempts to do just that through comparisons of sequenced genetic material. These researchers hope one day to better understand the origin of viruses—a discovery that could lead to advances in the treatments for the ailments they produce.
Viral Morphology
Viruses are noncellular, meaning they are biological entities that do not have a cellular structure. They therefore lack most of the components of cells, such as organelles, ribosomes, and the plasma membrane. A virion consists of a nucleic acid core, an outer protein coating or capsid, and sometimes an outer envelope made of protein and phospholipid membranes derived from the host cell. Viruses may also contain additional proteins, such as enzymes, within the capsid or attached to the viral genome. The most obvious difference between members of different viral families is the variation in their morphology, which is quite diverse. An interesting feature of viral complexity is that the complexity of the host does not necessarily correlate with the complexity of the virion. In fact, some of the most complex virion structures are found in the bacteriophages—viruses that infect the simplest living organisms, bacteria.
Morphology
Viruses come in many shapes and sizes, but these features are consistent for each viral family. As we have seen, all virions have a nucleic acid genome covered by a protective capsid. The proteins of the capsid are encoded in the viral genome, and are called capsomeres. Some viral capsids are simple helices or polyhedral “spheres,” whereas others are quite complex in structure (Figure 2).
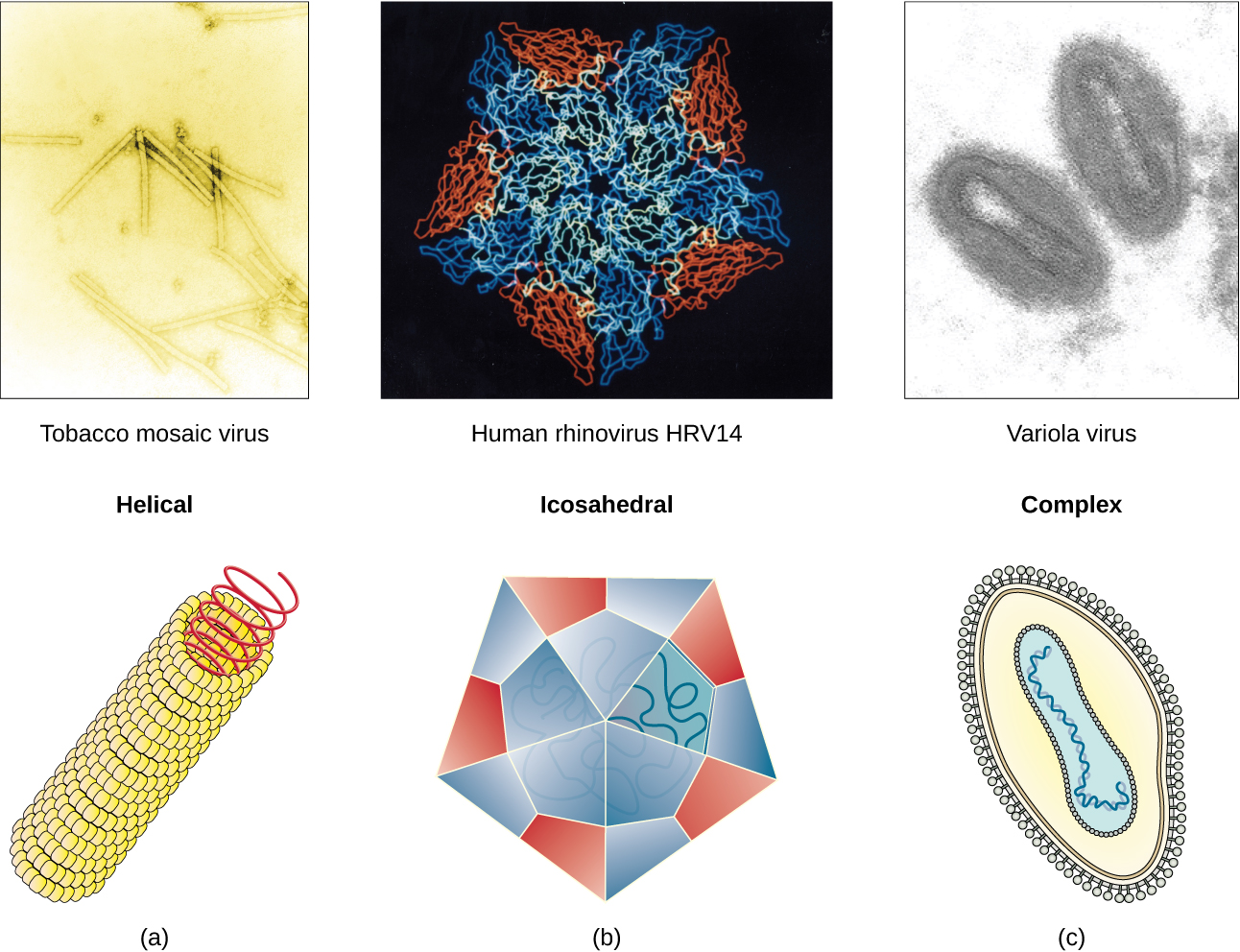
In general, the capsids of viruses are classified into four groups: helical, icosahedral, enveloped, and head-and-tail. Helical capsids are long and cylindrical. Many plant viruses are helical, including TMV. Icosahedral viruses have shapes that are roughly spherical, such as those of poliovirus or herpesviruses. Enveloped viruses have membranes derived from the host cell that surrounds the capsids. Animal viruses, such as HIV, are frequently enveloped. Head-and-tail viruses infect bacteria and have a head that is similar to icosahedral viruses and a tail shaped like helical viruses.
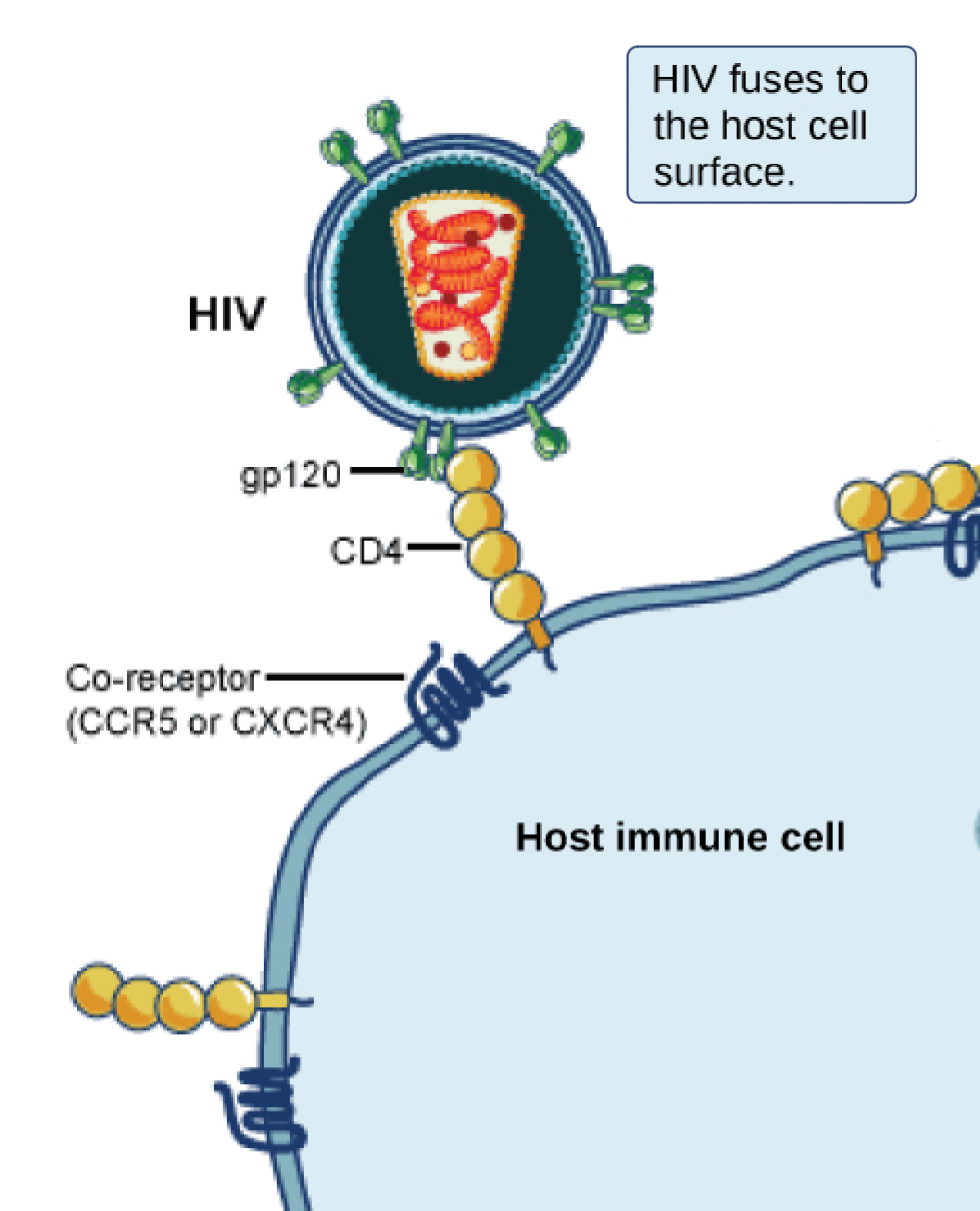
Many viruses use some sort of glycoprotein to attach to their host cells via molecules on the cell called viral receptors. For these viruses, attachment is required for later penetration of the cell membrane; only after penetration takes place can the virus complete its replication inside the cell. The receptors that viruses use are molecules that are normally found on cell surfaces and have their own physiological functions. It appears that viruses have simply evolved to make use of these molecules for their own replication. For example, HIV uses the CD4 molecule on T lymphocytes as one of its receptors (Figure 3). CD4 is a type of molecule called a cell adhesion molecule, which functions to keep different types of immune cells in close proximity to each other during the generation of a T lymphocyte immune response.
One of the most complex virions known, the T4 bacteriophage (which infects the Escherichia coli) bacterium, has a tail structure that the virus uses to attach to host cells and a head structure that houses its DNA.
Adenovirus, a non-enveloped animal virus that causes respiratory illnesses in humans, uses glycoprotein spikes protruding from its capsomeres to attach to host cells. Non-enveloped viruses also include those that cause polio (poliovirus), plantar warts (papillomavirus), and hepatitis A (hepatitis A virus).
Enveloped virions, such as the influenza virus, consist of nucleic acid (RNA in the case of influenza) and capsid proteins surrounded by a phospholipid bilayer envelope that contains virus-encoded proteins. Glycoproteins embedded in the viral envelope are used to attach to host cells. Other envelope proteins are the matrix proteins that stabilize the envelope and often play a role in the assembly of progeny virions. Chicken pox, HIV, and mumps are other examples of diseases caused by viruses with envelopes. Because of the fragility of the envelope, non-enveloped viruses are more resistant to changes in temperature, pH, and some disinfectants than enveloped viruses.
Overall, the shape of the virion and the presence or absence of an envelope tell us little about what disease the virus may cause or what species it might infect, but they are still useful means to begin viral classification (Figure 4).
VISUAL CONNECTION
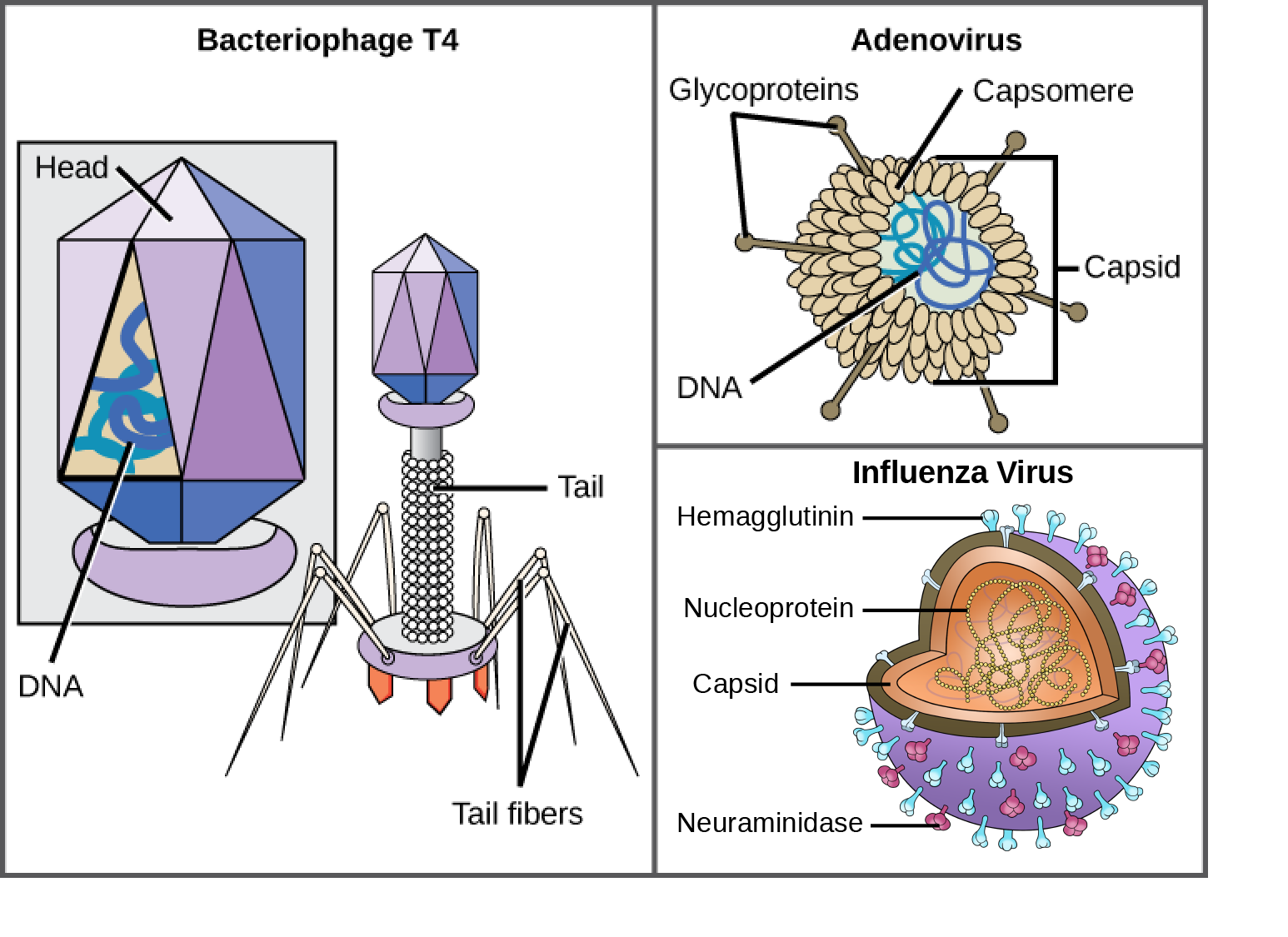
Which of the following statements about virus structure is true?
- All viruses are encased in a viral membrane.
- The capsomere is made up of small protein subunits called capsids.
- DNA is the genetic material in all viruses.
- Glycoproteins help the virus attach to the host cell.
Answer:
Statement “d” is true.
Types of Nucleic Acid
Unlike nearly all living organisms that use DNA as their genetic material, viruses may use either DNA or RNA. The virus core contains the genome—the total genetic content of the virus. Viral genomes tend to be small, containing only those genes that encode proteins which the virus cannot get from the host cell. This genetic material may be single- or double-stranded. It may also be linear or circular. While most viruses contain a single nucleic acid, others have genomes divided into several segments. The RNA genome of the influenza virus is segmented, which contributes to its variability and continuous evolution, and explains why it is difficult to develop a vaccine against it.
In DNA viruses, the viral DNA directs the host cell’s replication proteins to synthesize new copies of the viral genome and to transcribe and translate that genome into viral proteins. Human diseases caused by DNA viruses include chickenpox, hepatitis B, and adenoviruses. Sexually transmitted DNA viruses include the herpes virus and the human papilloma virus (HPV), which has been associated with cervical cancer and genital warts.
RNA viruses contain only RNA as their genetic material. To replicate their genomes in the host cell, the RNA viruses must encode their own enzymes that can replicate RNA into RNA or, in the retroviruses, into DNA. These RNA polymerase enzymes are more likely to make copying errors than DNA polymerases, and therefore often make mistakes during transcription. For this reason, mutations in RNA viruses occur more frequently than in DNA viruses. This causes them to change and adapt more rapidly to their host. Human diseases caused by RNA viruses include influenza, hepatitis C, measles, and rabies. The HIV virus, which is sexually transmitted, is an RNA retrovirus.
The Challenge of Virus Classification
Because most viruses probably evolved from different ancestors, the systematic methods that scientists have used to classify prokaryotic and eukaryotic cells are not very useful. If viruses represent “remnants” of different organisms, then even genomic or protein analysis is not useful. Why?, Because viruses have no common genomic sequence that they all share. For example, the 16S rRNA sequence so useful for constructing prokaryote phylogenies is of no use for a creature with no ribosomes! Biologists have used several classification systems in the past. Viruses were initially grouped by shared morphology. Later, groups of viruses were classified by the type of nucleic acid they contained, DNA or RNA, and whether their nucleic acid was single- or double-stranded. However, these earlier classification methods grouped viruses differently, because they were based on different sets of characters of the virus. The most commonly used classification method today is called the Baltimore classification scheme, and is based on how messenger RNA (mRNA) is generated in each particular type of virus.
Past Systems of Classification
Viruses contain only a few elements by which they can be classified: the viral genome, the type of capsid, and the envelope structure for the enveloped viruses. All of these elements have been used in the past for viral classification (Table 1 and Figure 5). Viral genomes may vary in the type of genetic material (DNA or RNA) and its organization (single- or double-stranded, linear or circular, and segmented or non-segmented). In some viruses, additional proteins needed for replication are associated directly with the genome or contained within the viral capsid.
| Genome Structure | Examples |
|---|---|
|
|
|
|
|
|
|
|
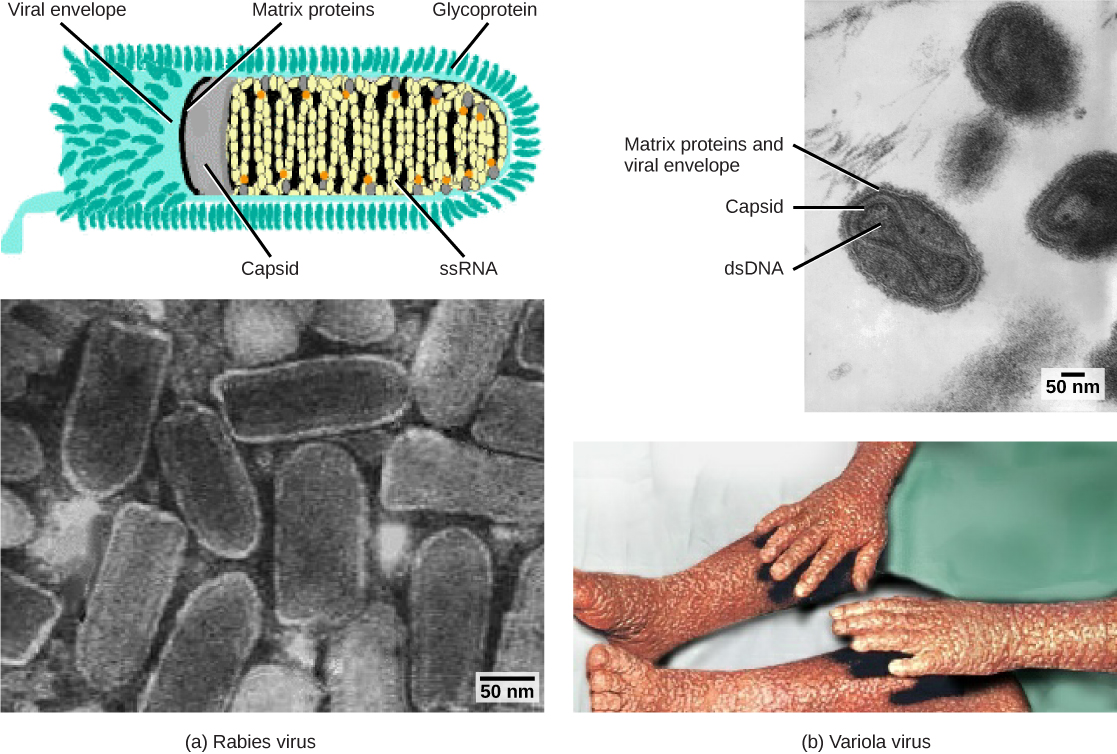
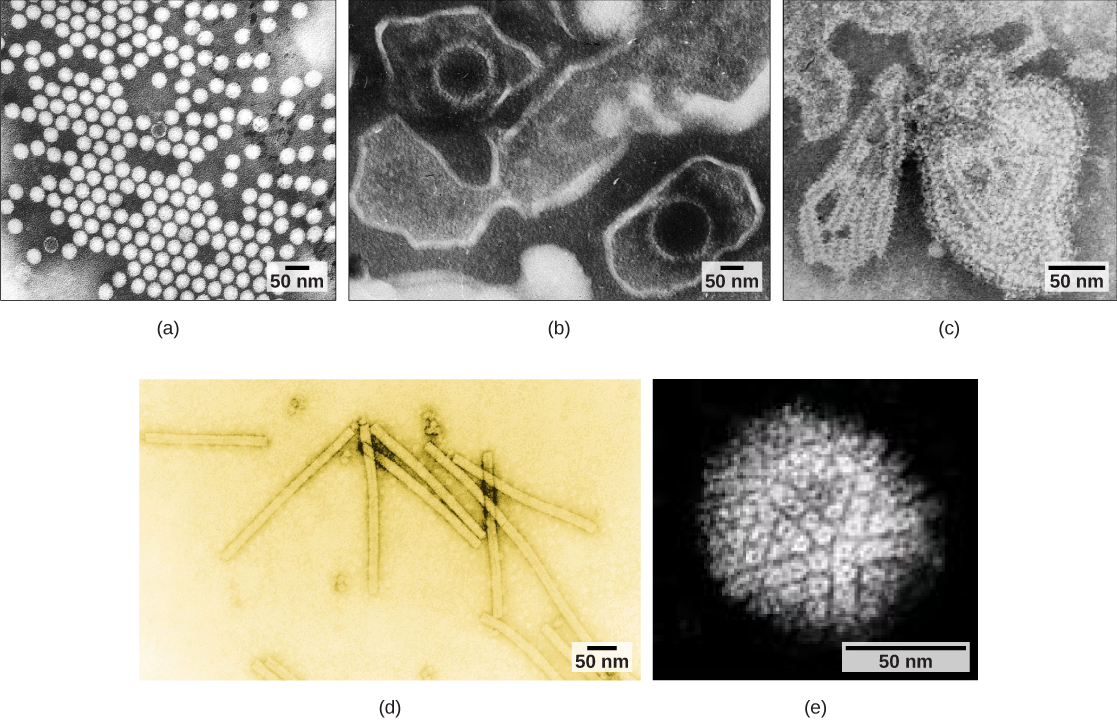
Viruses can also be classified by the design of their capsids (Table 2 and Figure 6). Capsids are classified as naked icosahedral, enveloped icosahedral, enveloped helical, naked helical, and complex. The type of genetic material (DNA or RNA) and its structure (single- or double-stranded, linear or circular, and segmented or non-segmented) are used to classify the virus core structures (Table 2)).
| Capsid Classification | Examples |
|---|---|
|
|
|
|
|
|
|
|
|
|
Baltimore Classification
The most commonly and currently used system of virus classification was first developed by Nobel Prize-winning biologist David Baltimore in the early 1970s. In addition to the differences in morphology and genetics mentioned above, the Baltimore classification scheme groups viruses according to how the mRNA is produced during the replicative cycle of the virus.
- Group I viruses contain double-stranded DNA (dsDNA) as their genome. Their mRNA is produced by transcription in much the same way as with cellular DNA, using the enzymes of the host cell.
- Group II viruses have single-stranded DNA (ssDNA) as their genome. They convert their single-stranded genomes into a dsDNA intermediate before transcription to mRNA can occur.
- Group III viruses use dsRNA as their genome. The strands separate, and one of them is used as a template for the generation of mRNA using the RNA-dependent RNA polymerase encoded by the virus.
- Group IV viruses have ssRNA as their genome with a positive polarity, which means that the genomic RNA can serve directly as mRNA. Intermediates of dsRNA, called replicative intermediates, are made in the process of copying the genomic RNA. Multiple, full-length RNA strands of negative polarity (complementary to the positive-stranded genomic RNA) are formed from these intermediates, which may then serve as templates for the production of RNA with positive polarity, including both full-length genomic RNA and shorter viral mRNAs.
- Group V viruses contain ssRNA genomes with a negative polarity, meaning that their sequence is complementary to the mRNA. As with Group IV viruses, dsRNA intermediates are used to make copies of the genome and produce mRNA. In this case, the negative-stranded genome can be converted directly to mRNA. Additionally, full-length positive RNA strands are made to serve as templates for the production of the negative-stranded genome.
- Group VI viruses have diploid (two copies) ssRNA genomes that must be converted, using the enzyme reverse transcriptase, to dsDNA; the dsDNA is then transported to the nucleus of the host cell and inserted into the host genome. Then, mRNA can be produced by transcription of the viral DNA that was integrated into the host genome.
- Group VII viruses have partial dsDNA genomes and make ssRNA intermediates that act as mRNA, but are also converted back into dsDNA genomes by reverse transcriptase, necessary for genome replication.
The characteristics of each group in the Baltimore classification are summarized in Table 3 with examples of each group.
| Group | Characteristics | Mode of mRNA Production | Example |
|---|---|---|---|
| I | Double-stranded DNA | mRNA is transcribed directly from the DNA template | Herpes simplex (herpesvirus) |
| II | Single-stranded DNA | DNA is converted to double-stranded form before RNA is transcribed | Canine parvovirus (parvovirus) |
| III | Double-stranded RNA | mRNA is transcribed from the RNA genome | Childhood gastroenteritis (rotavirus) |
| IV | Single stranded RNA (+) | Genome functions as mRNA | Common cold (picornavirus) |
| V | Single stranded RNA (-) | mRNA is transcribed from the RNA genome | Rabies (rhabdovirus) |
| VI | Single stranded RNA viruses with reverse transcriptase | Reverse transcriptase makes DNA from the RNA genome; DNA is then incorporated in the host genome; mRNA is transcribed from the incorporated DNA | Human immunodeficiency virus (HIV) |
| VII | Double stranded DNA viruses with reverse transcriptase | The viral genome is double-stranded DNA, but viral DNA is replicated through an RNA intermediate; the RNA may serve directly as mRNA or as a template to make mRNA | Hepatitis B virus (hepadnavirus) |
- acellular
- lacking cells
- capsid
- protein coating of the viral core
- capsomere
- protein subunit that makes up the capsid
- envelope
- lipid bilayer that encircles some viruses
- group I virus
- virus with a dsDNA genome
- group II virus
- virus with an ssDNA genome
- group III virus
- virus with a dsRNA genome
- group IV virus
- virus with an ssRNA genome with positive polarity
- group V virus
- virus with an ssRNA genome with negative polarity
- group VI virus
- virus with an ssRNA genome converted into dsDNA by reverse transcriptase
- group VII virus
- virus with a single-stranded mRNA converted into dsDNA for genome replication
- matrix protein
- envelope protein that stabilizes the envelope and often plays a role in the assembly of progeny virions
- negative polarity
- ssRNA viruses with genomes complementary to their mRNA
- positive polarity
- ssRNA virus with a genome that contains the same base sequences and codons found in their mRNA
- replicative intermediate
- dsRNA intermediate made in the process of copying genomic RNA
- reverse transcriptase
- enzyme found in Baltimore groups VI and VII that converts single-stranded RNA into double-stranded DNA
- viral receptor
- glycoprotein used to attach a virus to host cells via molecules on the cell
- virion
- individual virus particle outside a host cell
- virus core
- contains the virus genome